1.A.41 The Avian Reovirus p10 Viroporin/Plant Glutamine Dumper 1
(p10 Viroporin/GDU1) Family
Viroporins are alpha helical-type small proteins that are essential for normal viral function. Inhibitors have the potential of curing viral diseases. New-generation adamantane-based drugs are viroporin Iinhibitors, and one study focuses on SARS-CoV-2 viroporins. The detailed mechanism of these drugs against SARS-CoV-2 viroporin serves as a case study to guide future research on other viroporins. (Yousefbeigi and Marsusi 2025).
Structure-function traits common to all viroporins are their small sizes (ca. 60-120 aa), high hydrophobicities, and the presence of helical domains that transverse the membrane and assemble into oligomeric-permeating structures (Largo et al. 2016). The p10 protein is a membrane fusion/permeabilization protein encoded within the genomes of avian reoviruses. It is required for the virus to permeabilize cells during late infection and plays a role in virus-host interactions. Expression in bacterial cells arrests cell growth and enhances membrane permeability (Bodelón et al., 2002). The fusogenic extracellular N-terminal domain is not required for permeability. Therefore, it is a bifunctional protein where the two functions are associated with different domains. It is a small type I protein with 1 TMS (residues 43-63), its N-terminus out and its C-terminus in.
Novel pathways leading to disease stimulated by viroporin ion conduction, such as inflammasome driven immunopathology have been described (Nieto-Torres et al. 2015). A two stage folding/insertion mechanism for viroporins has been suggested (Martinez-Gil and Mingarro 2015). These proteins are crucial for the pathogenicity and replication of viruses as they aid in various stages of the viral life cycle, from genome uncoating to viral release. In addition, the ion channel activities of viroporins cause disruption of cellular ion (Ca2+) homeostasis. Fluctuation in the calcium level triggers activation of host defensive programmed cell death pathways as well as the inflammasome, which in turn are being subverted for the viruses' replication benefits (Sze and Tan 2015). The involvement of viroporins in virus-induced ER stress and autophagy has been discussed (Fung et al. 2015).
Glutamine Dumper 1. Phloem and xylem transport of amino acids involves two steps: export from one cell type to the apoplasm, and subsequent import into adjacent cells. High-affinity import is mediated by proton/amino acid cotransporters, while the mechanism of export remains unclear. Enhanced expression of the plant-specific type I membrane protein Glutamine Dumper1 (GDU1) has previously been shown to induce the secretion of glutamine from hydathodes and increased amino acid content in leaf apoplasm and xylem sap. Tolerance to low concentrations of amino acids and transport analyses demonstrated that net amino acid uptake is reduced in the glutamine-secreting GDU1 overexpressor gdu1-1D (Pratelli et al. 2010). The net uptake rate of phenylalanine decreased over time, and amino acid net efflux was increased in gdu1-1D compared with the wild type.
Independence of the export from proton gradients and ATP suggested that overexpression of GDU1 affects a passive export system. Each of the seven A. thaliana GDU genes led to similar phenotypes, including increased efflux of a wide spectrum of amino acids. Differences in expression profiles and functional properties suggested that the GDU genes fulfill different roles in roots, vasculature, and reproductive organs. Taken together, the GDUs appear to stimulate amino acid export by activating nonselective amino acid facilitators. GDU1 may either function as a channel or stimulate the activities of other exporters (Pratelli et al. 2010). It could be a subunit of an amino acid exporter because it seems to stimulate amino acid export by activating nonselective amino acid facilitators. The protein is required for the interaction with the RING-type E3 ubiquitin-protein ligase LOG2 to fulfill its function. It seems to play a role in the Gln export at hydathodes, at xylem parenchyma into xylem sap and from mesophyll into leaf apoplasm (Yu et al. 2015).
There are 2 subfamilies with TC#s 1.A.41.1 and 2. These subfamilies are either distantly related to each other, or are not related. Homology between these two 'subfamilies' has not been established.
The reaction catalyzed by p10 is:
Ions (in) 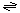 ions (out)
ions (out)