| 1.C.57 The Clostridial Cytotoxin (CCT) Family
Proteolytically processed clostridial cytotoxins A (306 kDa) and B (269 kDa) are O-glycosyltransferases that modify small GTPases of the Rho family by glucosylation of threonine residues, thereby blocking the action of the GTPases as switches of signal processes such as those mediated by the actin cytoskeleton. The toxins thus induce redistribution of actin filaments and cause the cells to round up. These cytotoxins are large (e.g., toxin B of C. difficile is 2366 aas long) and tripartite with the N-terminal domain being the catalytic unit, the C-terminal domain being the cellular receptor and the central hydrophobic domain being the channel-former. In this respect, they superficially resemble diphtheria toxin (BT; 1.C.7) although no significant sequence similarity between CCTs and BT is observed. The catalytic domains of CCTs probably enter the cytoplasm from acidic endosomes. The toxins form ion-permeable channels in cell membranes and artificial bilayers when exposed to acidic pH. pH-dependent channel formation has been demonstrated for C. difficile Toxin B and C as well as sordelli lethal toxin. Low pH presumably induces conformational/structural changes that promote membrane insertion and channel formation. Homologues are found in a variety of Gram-positive and Gram-negative bacteria. The E. coli toxin B protein and the Chlamydial TC0437 protein are of 3169 aas and 3255 aas, respectively. The distantly related ToxA toxin of Pasteurella multocida is 1285 aas while the E. coli Cnf1 and 2 toxins are 1014 aas, and the RTX cytotoxin of Vibrio vulnificus is 5206 aas.
Clostridium difficile toxins A and B are members of an important class
of virulence factors known as large clostridial toxins (LCTs). Toxin
action involves four major steps: receptor-mediated endocytosis,
translocation of a catalytic glucosyltransferase domain across the
membrane, release of the enzymatic moiety by autoproteolytic processing,
and a glucosyltransferase-dependent inactivation of Rho family
proteins. Pruitt et al. (2010) have imaged toxin A (TcdA) and toxin B (TcdB) holotoxins by
negative stain electron microscopy to show that these molecules are
similar in structure. They then determined a 3D structure for TcdA and
mapped the organization of its functional domains. The molecule has a
'pincher-like' head corresponding to the delivery domain and two tails,
long and short, corresponding to the receptor-binding and
glucosyltransferase domains, respectively. A second structure, obtained
at the acidic pH of an endosome, reveals a significant structural change
in the delivery and glucosyltransferase domains, and thus provides a
framework for understanding the molecular mechanism of LCT cellular
intoxication (Pruitt et al., 2010).
Clostridium difficile, the causative agent of nosocomial antibiotic-associated diarrhea and pseudomembranous colitis, possesses two main virulence factors: the large clostridial cytotoxins A and B. Cleavage of toxin B and all other large clostridial cytotoxins, is an autocatalytic process dependent on host cytosolic inositolphosphate cofactors. A covalent inhibitor of aspartate proteases, 1,2-epoxy-3-(p-nitrophenoxy)propane, completely blocks toxin B function on cultured cells and was used to identify the catalytically active protease site. The toxin uses eukaryotic signals for induced autoproteolysis to deliver its toxic domain into the cytosol of target cells. Reineke et al. (2007) present an integrated model for the uptake and inositolphosphate-induced activation of toxin B.
Clostridium difficile infection, caused by the actions of the homologous
toxins TcdA and TcdB on colonic epithelial cells is due to binding to target cells which triggers toxin
internalization into acidified vesicles, whereupon cryptic segments from within the 1,050-aa
translocation domain unfurl and insert into the bounding membrane, creating a transmembrane
passageway to the cytosol (Zhang et al. 2014). Sensitive residues-clustered between amino acyl residues 1,035
and 1,107, when individually mutated, reduced cellular toxicity by >1,000-fold. Defective variants exhibit impaired pore formation in planar lipid bilayers
and biological membranes, resulting in an inability to intoxicate cells through either apoptotic or
necrotic pathways. The findings suggest similarities between the pore-
forming 'hotspots' of TcdB and the diphtheria toxin translocation
domain (Zhang et al. 2014).
The generalized transport reactions catalyzed by CCTs are:
N-terminal catalytic domain (out) → N-terminal catalytic domain (in)
Ions and other solutes (in) 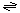 ions and other solutes (out ions and other solutes (out
|
This family belongs to the RTX-toxin Superfamily.
|
| References: |
Amimoto, K., T. Noro, E. Oishi, and M. Shimizu. (2007). A novel toxin homologous to large clostridial cytotoxins found in culture supernatant of Clostridium perfringens type C. Microbiology. 153: 1198-1206.
|
Baldwin, M.R., J.H. Lakey, and A.J. Lax. (2004). Identification and characterization of the Pasteurella multocida toxin translocation domain. Mol. Microbiol. 54: 239-250.
|
Barth, H., G. Pfeifer, F. Hofmann, E. Maier, R. Benz, and K. Aktories. (2001). Low pH-induced formation of ion channels by Clostridium difficile toxin B in target cells. J. Biol. Chem. 276: 10670-10676.
|
Belland, R.J., M.A. Scidmore, D.D. Crane, D.M. Hogan, W. Whitmire, G. McClarty, and H.D. Caldwell. (2001). Chlamydia trachomatis cytotoxicity associated with complete and partial cytotoxin genes. Proc. Natl. Acad. Sci. USA 98: 13984-13989.
|
Genisyuerek, S., P. Papatheodorou, G. Guttenberg, R. Schubert, R. Benz, and K. Aktories. (2011). Structural determinants for membrane insertion, pore formation and translocation of Clostridium difficile toxin B. Mol. Microbiol. 79: 1643-1654.
|
Hamza, T., Z. Zhang, R.A. Melnyk, and H. Feng. (2016). Defective mutations within the translocation domain of Clostridium difficile toxin B impair disease pathogenesis. Pathog Dis 74: ftv098.
|
Oswald, E., M. Sugai, A. Labigne, H.C. Wu, C. Fiorentini, P. Boquet, and A.D. O'Brien. (1994). Cytotoxic necrotizing factor type 2 produced by virulent Escherichia coli modifies the small GTP-binding proteins Rho involved in assembly of actin stress fibers. Proc. Natl. Acad. Sci. USA 91: 3814-3818.
|
Pruitt, R.N., M.G. Chambers, K.K. Ng, M.D. Ohi, and D.B. Lacy. (2010). Structural organization of the functional domains of Clostridium difficile toxins A and B. Proc. Natl. Acad. Sci. USA 107: 13467-13472.
|
Reineke, J., S. Tenzer, M. Rupnik, A. Koschinski, O. Hasselmayer, A. Schrattenholz, H. Schild, and C. von Eichel-Streiber. (2007). Autocatalytic cleavage of Clostridium difficile toxin B. Nature 446: 415-419.
|
Satchell, K.J.F. (2015). Multifunctional-autoprocessing repeats-in-toxin (MARTX) Toxins of Vibrios. Microbiol Spectr 3:.
|
Sheahan, K.L., C.L. Cordero, and K.J. Satchell. (2007). Autoprocessing of the Vibrio cholerae RTX toxin by the cysteine protease domain. EMBO. J. 26: 2552-2561.
|
Shen, A., P.J. Lupardus, V.E. Albrow, A. Guzzetta, J.C. Powers, K.C. Garcia, and M. Bogyo. (2009). Mechanistic and structural insights into the proteolytic activation of Vibrio cholerae MARTX toxin. Nat Chem Biol 5: 469-478.
|
Woida, P.J. and K.J.F. Satchell. (2018). Coordinated delivery and function of bacterial MARTX toxin effectors. Mol. Microbiol. 107: 133-141.
|
Zhang Z., Park M., Tam J., Auger A., Beilhartz GL., Lacy DB. and Melnyk RA. (2014). Translocation domain mutations affecting cellular toxicity identify the Clostridium difficile toxin B pore. Proc Natl Acad Sci U S A. 111(10):3721-6.
|
Zhao, J.F., A.H. Sun, P. Ruan, X.H. Zhao, M.Q. Lu, and J. Yan. (2009). Vibrio vulnificus cytolysin induces apoptosis in HUVEC, SGC-7901 and SMMC-7721 cells via caspase-9/3-dependent pathway. Microb. Pathog. 46: 194-200.
|
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.C.57.1.1 | Cytotoxin B, TcdB. The minimal pore-forming region is within amino acid residues 830 and 990 including glutamate-970 and glutamate-976. These two residues are essential for pore formation (Genisyuerek et al., 2011). Other residues important for toxicity have been identified (Zhang et al. 2014). Residues in the translocation domain of TcdB that form the pore and function in toxin translocation have been identified (Hamza et al. 2016). | Bacteria | Cytotoxin B (TcdB) of Clostridium difficile |
| |
| 1.C.57.1.2 | Cytotoxin A | Bacteria | Cytotoxin A of Clostridium difficile |
| |
| 1.C.57.1.3 | Lethal toxin | Bacteria | Lethal toxin (cytotoxin L) of Clostridium sordellii |
| |
| 1.C.57.1.4 | α-toxin | Bacteria | α-toxin of Clostridium novyi |
| |
| 1.C.57.1.5 | Cytotoxin C, TpeL (Amimoto et al., 2007) | Bacteria | TpeL of Clostridium difficile (A2PYQ6) |
| |
| 1.C.57.1.6 | MCF toxin of 2993 aas | Proteobacteria | MCF toxin of Photorhabdus asymbiotica subsp. asymbiotica (Xenorhabdus luminescens) |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.C.57.2.1 | Toxin B | Bacteria | Toxin B of E. coli plasmid p0157 |
| |
| 1.C.57.2.2 | Cytotoxic adherence factor TC0437 | Bacteria | TC0437 of Chlamydia muridarum |
| |
| 1.C.57.2.3 | LifA/Efa1-related large cytotoxin of 3218 aas. | | LifA of Chlamydia muridarum |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.C.57.3.1 | Pasteurella multocida toxin (PMT); dermonecrotic toxin (DMT); mitogenic toxin (ToxA) (Baldwin et al., 2004) | Bacteria | PMT of Pasteurella multocida (P17452) |
| |
| 1.C.57.3.2 | Cytotoxic necrotizing factor type 1, Cnf1 (Oswald et al., 1994) | Bacteria | Cnf1 of E. coli (AAN03786) |
| |
| 1.C.57.3.3 | Cytotoxic necrotizing factor type 2, Cnf2 (Oswald et al., 1994) | Bacteria | Cnf2 of E. coli (A55260) |
| |
| 1.C.57.3.4 | RTX (repeat in toxin) cytotoxin of 5206 aas, also called the "multifunctional-autoprocessing RTX" (MARTXVv) toxin, or Vibrio vulnificus cytotoxin (VVC). It exists in at least four distinct variants of the rtxA1 gene that encode toxins with different arrangements of effector domains that arose by recombination. VVC, in addition to being a pore-forming toxin, may be a transmembrane toxin with the ability to induce
apoptosis in human vascular endothelial cells and tumor cells (Zhao et al. 2009). The protein has an α,β-hydrolase domain (residues ~2900 - 3120). | Bacteria | RTX cytotoxin of Vibrio vulnificus (BAC97056) |
| |
| 1.C.57.3.5 | Multifunctional-autoprocessing repeats-in-toxin, RtxA or Rtx, of 4558 aas. It is the precursor
of a multifunctional toxin that causes destruction of the actin
cytoskeleton by covalent cross-linking of actin and inactivation of Rho
GTPases when translocated into the host cytoplasm (Satchell 2015). Upon translocation into the host cell, it undergoes autoprocessing in cis
mediated by the peptidase C80 domain (also named CPD domain). The
protease activity is activated upon binding inositol hexakisphosphate
(InsP6), present at the host cell membrane, delivering the cysteine
protease domain-containing toxin F3 chain to the host cytosol (Sheahan et al. 2007, Shen et al. 2009). It forms a pore in the plasma membrane of a eukaryotic cell to deliver the toxin (Woida and Satchell 2018). | | RtxA of Vibrio cholerae |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.C.57.4.1 | Putative toxin A of 294 aas | Spirochaetes | Toxin A of Brachyspira intermedia |
| |
 ions and other solutes (out
ions and other solutes (out