3.D.9 The H+-translocating F420H2 Dehydrogenase (F420H2DH) Family
A single F420H2 dehydrogenase (also referred to as F420H2: quinol oxidoreductase) from the methanogenic archaeon, Methanosarcina mazei Gö1, has been shown to be a redox driven proton pump. Reduction of 2-hydroxyphenazine by F420H2DH is accompanied by the translocation of 1 H+ per 2 electrons transferred. The gene cluster encoding the F420H2DH includes 12 genes, fpoABCDHIJKLMNO. Several of the subunits are related to those of the mitochondrial 'complex I' NDH family members (TC#3.D.1). Thus, the gene products, FpoA, H, J, K, L, M and N, are highly hydrophobic and are homologous to subunits that form the membrane integral module of NDH-1. FpoB, C, D and I have their counterparts in the amphipathic membrane-associated module of NDH-1. However, homologues of the hydrophilic NADH-oxidizing input are absent. Instead, FpoF probably catalyzes F420H2 oxidation, providing the input module for transferring electrons to the membrane module, while one or more subunits (the output module) may be adapted to the reduction of methanophenazine.
The F420H2DH of M. mazei has a molecular size of about 120 kDa and contains Fe-S clusters and FAD. A similar 5 subunit enzyme has been isolated from Methanolobus tindarius. The sulfate-reducing Archaeoglobus fulgidus (and several other archaea) also have this enzyme.
Methanomassiliicoccus luminyensis has been isolated from the human gut and requires H2 and methanol or methylamines to produce methane. The organism lacks cytochromes indicating that it cannot couple membrane-bound electron transfer reactions with the extrusion of protons or sodium ions using known methanogenic pathways. Furthermore, M. luminyensis contains a soluble MvhAGD/HdrABC complex as found in obligate hydrogenotrophic methanogens but the energy conserving methyltransferase (MtrA-H) is absent. Kröninger et al. 2015 presented evidence that M. luminyensis uses two types of heterodisulfide reductases (HdrABC and HdrD) in an energy conserving process. RT-qPCR studies revealed that genes coding for both heterodisulfide reductases showed high expression levels. Other genes with high transcript abundance were fpoA as part of the operon encoding the 'headless' F420 H2 dehydrogenase and atpB as part of the operon encoding the A1 Ao ATP synthase. High activities of the soluble heterodisulfide reductase HdrABC and the hydrogenase MvhADG were found in the cytoplasm. Also, heterologously produced HdrD could reduce CoM-S-S-CoB using reduced methylviologen as electron donor. Kröninger et al. 2015 proposed that membrane-bound electron transfer is based on the conversion of two molecules of methanol and the concurrent formation of two molecules of the heterodisulfide CoM-S-S-CoB. First the HdrABC/MvhADG complex catalyzes the H2-dependent reduction of CoM-S-S-CoB and the formation of reduced ferredoxin. In a second cycle reduced ferredoxin is oxidized by the 'headless' F420 H2 dehydrogenase thereby translocating up to 4H+ across the membrane, and electrons are channeled to HdrD for the reduction of the second heterodisulfide.
The overall vectorial reaction catalyzed by F420H2DH is:
Reduced donor (2e-) + nH+ (in) 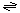 oxidized acceptor (2e-) + nH+ (out)
oxidized acceptor (2e-) + nH+ (out)