| 1.B.1 The General Bacterial Porin (GBP) Family
OMP porins are present in the outer membranes of Gram-negative bacteria, mitochondria and plastids. They catalyze the energy-independent facilitation of small (Mr of <1000 Da) molecules across the outer membranes of bacteria and organelles with variable degrees of selectivity. The structurally characterized members of this functional superfamily usually consist of homotrimeric proteins with subunits that are of 250-450 amino acyl residues in length. The high resolution three-dimensional structures of several of these proteins are known. These proteins include OmpC, OmpF and PhoP of E. coli. They form 16-stranded antiparallel β-barrel structures with all β-strands hydrogen-bonded to their nearest neighbors along the chain. Each trimer consists of three channels, each with the β-barrel perpendicular to the plane of the membrane. Polypeptide loops lining the inner barrel wall restrict the channel width, thereby defining the diffusion properties of the pore. Some porins are cation-selective, others are anion-selective and still others are selective for specific compounds (e.g., sugars, nucleotides, phosphate, pyrophosphate). Sequence comparisons and three-dimensional structural analyses suggest that many of the families described under category 1.B are related (see porin superfamilies in TCDB and (Reddy and Saier 2016). Outer membrane porins have been reviewed (Masi et al. 2019; Vergalli et al. 2019). Hermansen et al. 2022 have summarized the knowledge of beta-barrel structure and folding and give an overview of their functions, evolution, and potential as drug targets. Larger porins can be made from smaller porins using a loop-to-hairpin mechanism (Dhar et al. 2023). Gating of beta-barrel protein pores, porins and channels has been reviewed (Mayse and Movileanu 2023). Structural bioinformatic studies of bacterial outer membrane beta-barrel porins have been conducted (Sajeev-Sheeja et al. 2023). Porin permeabilities and minimal inhibitory concentrations have been predicted through machine learning (Boi et al. 2025).
β-barrel membrane proteins perform a variety of functions, such as mediating non-specific, passive transport of ions and small molecules, selectively passing molecules like maltose and sucrose, and can form voltage dependent anion channels. Understanding the structural features of β-barrel membrane proteins and detecting them in genomic sequences are challenging tasks in structural and functional genomics. With the survey of experimentally known amino acid sequences and structures, the characteristic features of amino acid residues in β-barrel membrane proteins and novel parameters for understanding their folding and stability have been described by Gromiha and Suwa (2007). Statistical methods and machine learning techniques discriminate β-barrel membrane proteins from other folding types of globular and membrane proteins. Different methods including hydrophobicity profiles, rule based approach, amino acid properties, neural networks, hidden Markov models etc., predict membrane spanning segments of β-barrel membrane proteins. Discrimination techniques for detecting β-barrel membrane proteins in genomic sequences are discussed by Gromiha and Suwa (2007).
Vibrio furnissii possesses an outer membrane porin that is induced by β1,4-N-acetyl glucosamine (GlcNAc) oligomers of two to six sugar units, hydrolysis products of chitinase action on chitin (Keyhani et al., 2000). This porin is required for growth on (GlcNAc)3, and it transports acetylated chitobiose analogues, suggesting that it is specific for these oligosaccharides. It forms a subfamily (TC #1.B.1.7.1) of the GBP family. Another porin, OmpP2 of Haemophilus influenzae (TC # 1.B.1.3.2), shows specificity for nicotinamide-derived nucleotide substrates (Andersen et al., 2003).
Gram-negative Legionella pneumophila produces a siderophore (legiobactin) that promotes lung infection. lbtA and lbtB are required for the synthesis and secretion of legiobactin. An iron-repressed gene (lbtU) is directly upstream of the lbtAB-containing operon. LbtU is an outer membrane protein consisting of a 16-stranded transmembrane β-barrel, multiple extracellular domains, and short periplasmic tails. Although replicating normally, lbtU mutants, like lbtA mutants, were impaired for growth on iron-depleted media and would not take up Fe3+ legiobactin. It is the Legionella siderophore receptor.
The generalized transport reaction catalyzed by porins is:
Solute (out) 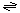 Solute (in) Solute (in)
|
This family belongs to the Outer Membrane Pore-forming Protein I (OMPP-I) Superfamily .
|
| References: |
Acharya, A., P.K. Behera, and U. Kleinekathöfer. (2024). Molecular Mechanism of Ciprofloxacin Translocation Through the Major Diffusion Channels of the ESKAPE Pathogens and. J Phys Chem B 128: 8376-8387.
|
Andersen, C., E. Maier, G. Kemmer, J. Blass, A.-K. Hilpert, R. Benz, and J. Reidl. (2003). Porin OmpP2 of Haemophilus influenzae shows specificity for nicotinamide-derived nucleotide substrates. J. Biol. Chem. 278: 24269-24276.
|
Andrade, I.d.a.S., J.L. Vianez-Júnior, C.L. Goulart, F. Homblé, J.M. Ruysschaert, W.M. Almeida von Krüger, P.M. Bisch, W. de Souza, R. Mohana-Borges, and M.C. Motta. (2011). Characterization of a porin channel in the endosymbiont of the trypanosomatid protozoan Crithidia deanei. Microbiology 157: 2818-2830.
|
Aunkham, A., A. Schulte, M. Winterhalter, and W. Suginta. (2014). Porin involvement in cephalosporin and carbapenem resistance of Burkholderia pseudomallei. PLoS One 9: e95918.
|
Aunkham, A., A. Schulte, W.C. Sim, W. Chumjan, and W. Suginta. (2020). Vibrio campbellii chitoporin: Thermostability study and implications for the development of therapeutic agents against Vibrio infections. Int J Biol Macromol. [Epub: Ahead of Print]
|
Aunkham, A., M. Zahn, A. Kesireddy, K.R. Pothula, A. Schulte, A. Baslé, U. Kleinekathöfer, W. Suginta, and B. van den Berg. (2018). Structural basis for chitin acquisition by marine Vibrio species. Nat Commun 9: 220.
|
Bajaj, H., Q.T. Tran, K.R. Mahendran, C. Nasrallah, J.P. Colletier, A. Davin-Regli, J.M. Bolla, J.M. Pagès, and M. Winterhalter. (2012). Antibiotic uptake through membrane channels: role of Providencia stuartii OmpPst1 porin in carbapenem resistance. Biochemistry 51: 10244-10249.
|
Bartsch, A., S. Llabrés, F. Pein, C. Kattner, M. Schön, M. Diehn, M. Tanabe, A. Munk, U. Zachariae, and C. Steinem. (2019). High-resolution experimental and computational electrophysiology reveals weak β-lactam binding events in the porin PorB. Sci Rep 9: 1264.
|
Belousov, M.V., A.O. Kosolapova, H. Fayoud, M.I. Sulatsky, A.I. Sulatskaya, M.N. Romanenko, A.G. Bobylev, K.S. Antonets, and A.A. Nizhnikov. (2023). OmpC and OmpF Outer Membrane Proteins of and Form Amyloids. Int J Mol Sci 24:.
|
Benz, R., R.P. Darveau, and R.E.W. Hancock. (1984). Outer-membrane protein PhoE from Escherichia coli forms anion-selective pores in lipid-bilayer membranes. Eur. J. Biochem. 140: 319-324.
|
Blasband, A.J. and C.A. Schnaitman. (1987). Regulation in Escherichia coli of the porin protein gene encoded by lambdoid bacteriophages. J. Bacteriol. 169: 2171-2176.
|
Boi, S., S. Puxeddu, I. Delogu, D. Farci, D. Piano, A. Manzin, M. Ceccarelli, F. Angius, M.A. Scorciapino, and S. Milenkovic. (2025). Seeking Correlation Among Porin Permeabilities and Minimum Inhibitory Concentrations Through Machine Learning: A Promising Route to the Essential Molecular Descriptors. Molecules 30:.
|
Bornet, C., N. Saint, L. Fetnaci, M. Dupont, A. Davin-Régli, C. Bollet, and J.M. Pagès. (2004). Omp35, a new Enterobacter aerogenes porin involved in selective susceptibility to cephalosporins. Antimicrob. Agents Chemother. 48: 2153-2158.
|
Brunen, M., H. Engelhardt, A. Schmid, and R. Benz. (1991). The major outer membrane protein of Acidovorax delafieldii is an anion-selective porin. J. Bacteriol. 173: 4182-4187.
|
Brunson, D.N., E. Maldosevic, A. Velez, E. Figgins, and T.N. Ellis. (2019). Porin loss in Klebsiella pneumoniae clinical isolates impacts production of virulence factors and survival within macrophages. Int. J. Med. Microbiol. 309: 213-224.
|
Calderón, I.L., E. Morales, N.J. Caro, C.A. Chahúan, B. Collao, F. Gil, J.M. Villarreal, F. Ipinza, G.C. Mora, and C.P. Saavedra. (2010). Response regulator ArcA of Salmonella enterica serovar Typhimurium downregulates expression of OmpD, a porin facilitating uptake of hydrogen peroxide. Res. Microbiol. 162: 214-222.
|
Cervera, J., A.G. Komarov, and V.M. Aguilella. (2008). Rectification properties and pH-dependent selectivity of meningococcal class 1 porin. Biophys. J. 94: 1194-1202.
|
Chang HK., Dennis JJ. and Zylstra GJ. (2009). Involvement of two transport systems and a specific porin in the uptake of phthalate by Burkholderia spp. J Bacteriol. 191(14):4671-3.
|
Chatfield, C.H., B.J. Mulhern, D.M. Burnside, and N.P. Cianciotto. (2011). Legionella pneumophila LbtU acts as a novel, TonB-independent receptor for the legiobactin siderophore. J. Bacteriol. 193: 1563-1575.
|
Chistyulin, D.K., O.D. Novikova, E.A. Zelepuga, V.A. Khomenko, G.N. Likhatskaya, O.Y. Portnyagina, and Y.N. Antonenko. (2019). An Abnormally High Closing Potential of the OMPF Porin Channel from Yersinia Ruckeri: The Role of Charged Residues and Intramolecular Bonds. Acta Naturae 11: 89-98.
|
Chumjan W., Winterhalter M., Schulte A., Benz R. and Suginta W. (2015). Chitoporin from the Marine Bacterium Vibrio harveyi: PROBING THE ESSENTIAL ROLES OF TRP136 AT THE SURFACE OF THE CONSTRICTION ZONE. J Biol Chem. 290(31):19184-96.
|
Chumjan, W., M. Winterhalter, and W. Suginta. (2019). Effects of H-bonds on sugar binding to chitoporin from Vibrio harveyi. Biochim. Biophys. Acta. Biomembr 1861: 610-618.
|
Cowan, S.W., T. Schirmer, G. Rummel, M. Steiert, R. Ghosh, R.A. Pauptit, J.N. Jansonius, and J.P. Rosenbusch. (1992). Crystal structures explain functional properties of two E. coli porins. Nature 358: 727-733.
|
Dam, S., J.M. Pagès, and M. Masi. (2017). Dual Regulation of the Small RNA MicC and the Quiescent Porin OmpN in Response to Antibiotic Stress in Escherichia coli. Antibiotics (Basel) 6:.
|
Dé, E., A. Baslé, M. Jaquinod, N. Saint, M. Malléa, G. Molle, and J.M. Pagès. (2001). A new mechanism of antibiotic resistance in Enterobacteriaceae induced by a structural modification of the major porin. Mol. Microbiol. 41: 189-198.
|
Delcour, A.H. (1997). Function and modulation of bacterial porins: insights from electrophysiology. FEMS Microbiol. Lett. 152: 115-123.
|
Dhar, V., S. Gandhi, S.C. Sakharwade, A. Chawla, and A. Mukhopadhaya. (2023). Vibrio cholerae Porin OmpU Activates Dendritic Cells via TLR2 and the NLRP3 Inflammasome. Infect. Immun. 91: e0033222.
|
Donoghue, A., M. Winterhalter, and T. Gutsmann. (2023). Influence of Membrane Asymmetry on OmpF Insertion, Orientation and Function. Membranes (Basel) 13:.
|
Duperthuy, M., A.E. Sjöström, D. Sabharwal, F. Damghani, B.E. Uhlin, and S.N. Wai. (2013). Role of the Vibrio cholerae matrix protein Bap1 in cross-resistance to antimicrobial peptides. PLoS Pathog 9: e1003620.
|
Duret, G. and A.H. Delcour. (2010). Size and dynamics of the Vibrio cholerae porins OmpU and OmpT probed by polymer exclusion. Biophys. J. 98: 1820-1829.
|
Easton, D.M., A. Smith, S.G. Gallego, A.R. Foxwell, A.W. Cripps, and J.M. Kyd. (2005). Characterization of a novel porin protein from Moraxella catarrhalis and identification of an immunodominant surface loop. J. Bacteriol. 187: 6528-6535.
|
Egger, L.A., H. Park, and M. Inouye. (1997). Signal transduction via the histidyl-aspartyl phosphorelay. Genes Cells 2: 167-184.
|
El-Khatib, M., C. Nasrallah, J. Lopes, Q.T. Tran, G. Tetreau, H. Basbous, D. Fenel, B. Gallet, M. Lethier, J.M. Bolla, J.M. Pagès, M. Vivaudou, M. Weik, M. Winterhalter, and J.P. Colletier. (2018). Porin self-association enables cell-to-cell contact in floating communities. Proc. Natl. Acad. Sci. USA 115: E2220-E2228.
|
Elazar, M., D. Halfon, I. Pechatnikov, and Y. Nitzan. (2007). Porin isolated from the outer membrane of Erwinia amylovora and its encoding gene. Curr. Microbiol. 54: 155-161.
|
Fàbrega, A., J.L. Rosner, R.G. Martin, M. Solé, and J. Vila. (2012). SoxS-dependent coregulation of ompN and ydbK in a multidrug-resistant Escherichia coli strain. FEMS Microbiol. Lett. 332: 61-67.
|
Fernández-Mora, M., J.L. Puente, and E. Calva. (2004). OmpR and LeuO positively regulate the Salmonella enterica serovar Typhi ompS2 porin gene. J. Bacteriol. 186: 2909-2920.
|
Forst, S., J. Waukau, G. Leisman, M. Exner, and R. Hancock. (1995). Functional and regulatory analysis of the OmpF-like porin, OpnP, of the symbiotic bacterium Xenorhabdus nematophilus. Mol. Microbiol. 18: 779-789.
|
Forst, S.A. and N. Tabatabai. (1997). Role of the histidine kinase, EnvZ, in the production of outer membrane proteins in the symbiotic-pathogenic bacterium Xenorhabdus nematophilus. Appl. Environ. Microbiol. 63: 962-968.
|
Fralick, J.A., D.L. Diedrich, and S. Casey-Wood. (1990). Isolation of an Lc-specific Escherichia coli bacteriophage. J. Bacteriol. 172: 1660-1662.
|
Ganguly B., Tewari K. and Singh R. (2015). Homology modeling, functional annotation and comparative genomics of outer membrane protein H of Pasteurella multocida. J Theor Biol. 386:18-24.
|
Gao, T., L. Ju, J. Yin, and H. Gao. (2015). Positive regulation of the Shewanella oneidensis OmpS38, a major porin facilitating anaerobic respiration, by Crp and Fur. Sci Rep 5: 14263.
|
Gao, Y., G. Widmalm, and W. Im. (2025). Dynamics and Interactions of OmpF Porin in an Asymmetric Bacterial Outer Membrane including LPS, ECA, and CPS. Biomacromolecules 26: 3711-3720.
|
García-Sureda, L., A. Doménech-Sánchez, M. Barbier, C. Juan, J. Gascó, and S. Albertí. (2011). OmpK26, a novel porin associated with carbapenem resistance in Klebsiella pneumoniae. Antimicrob. Agents Chemother. 55: 4742-4747.
|
Gensberg, K., A.W. Smith, F.S. Brinkman, and R.E. Hancock. (1999). Identification of oprG, a gene encoding a major outer membrane protein of Pseudomonas aeruginosa. J. Antimicrob. Chemother. 43: 607-608.
|
Ghale G., Lanctot AG., Kreissl HT., Jacob MH., Weingart H., Winterhalter M. and Nau WM. (2014). Chemosensing ensembles for monitoring biomembrane transport in real time. Angew Chem Int Ed Engl. 53(10):2762-5.
|
Golla, V.K., E. Sans-Serramitjana, K.R. Pothula, L. Benier, J.A. Bafna, M. Winterhalter, and U. Kleinekathöfer. (2019). Fosfomycin Permeation through the Outer Membrane Porin OmpF. Biophys. J. 116: 258-269.
|
Goulart, C.L., P.M. Bisch, W.M. von Krüger, and F. Homblé. (2015). VCA1008: An Anion-Selective Porin of Vibrio Cholerae. Biochim. Biophys. Acta. 1848: 680-687.
|
Grant, T.A., M. López-Pérez, J.M. Haro-Moreno, and S. Almagro-Moreno. (2023). Allelic diversity uncovers protein domains contributing to the emergence of antimicrobial resistance. PLoS Genet 19: e1010490.
|
Gromiha, M.M., and M. Suwa (2007). Current developments on β- barrel membrane proteins: sequence and structure analysis, discrimination and prediction. Curr. Protein Pept. Sci. 8: 580-599.
|
Gulati, A., R. Kumar, and A. Mukhopadhaya. (2019). Differential Recognition of OmpU by Toll-Like Receptors in Monocytes and Macrophages for the Induction of Proinflammatory Responses. Infect. Immun. 87:.
|
Gupta S., Prasad GV. and Mukhopadhaya A. (2015). Vibrio cholerae Porin OmpU Induces Caspase-independent Programmed Cell Death upon Translocation to the Host Cell Mitochondria. J Biol Chem. 290(52):31051-68.
|
Hadi-Alijanvand, H. and M. Rouhani. (2015). Journey of Poly-Nucleotides through OmpF Porin. J Phys Chem B 119: 6113-6128.
|
Hamzaoui, Z., A. Ocampo-Sosa, M.F. Martinez, S. Landolsi, S. Ferjani, E. Maamar, M. Saidani, A. Slim, L. Martinez-Martinez, and I.B. Boubaker. (2018). Role of association of OmpK35 and OmpK36 alteration and bla and/or bla in conferring carbapenem resistance among non-producing carbapenemase-Klebsiella pneumoniae. Int J Antimicrob Agents. [Epub: Ahead of Print]
|
Hermansen, S., D. Linke, and J.C. Leo. (2022). Transmembrane β-barrel proteins of bacteria: From structure to function. Adv Protein Chem Struct Biol 128: 113-161.
|
Housden, N.G., J.A. Wojdyla, J. Korczynska, I. Grishkovskaya, N. Kirkpatrick, A.M. Brzozowski, and C. Kleanthous. (2010). Directed epitope delivery across the Escherichia coli outer membrane through the porin OmpF. Proc. Natl. Acad. Sci. USA 107: 21412-21417.
|
Housden, N.G., J.T. Hopper, N. Lukoyanova, D. Rodriguez-Larrea, J.A. Wojdyla, A. Klein, R. Kaminska, H. Bayley, H.R. Saibil, C.V. Robinson, and C. Kleanthous. (2013). Intrinsically disordered protein threads through the bacterial outer-membrane porin OmpF. Science 340: 1570-1574.
|
Housden, N.G., P. Rassam, S. Lee, F. Samsudin, R. Kaminska, C. Sharp, J.D. Goult, M.L. Francis, S. Khalid, H. Bayley, and C. Kleanthous. (2018). Directional Porin Binding of Intrinsically Disordered Protein Sequences Promotes Colicin Epitope Display in the Bacterial Periplasm. Biochemistry 57: 4374-4381.
|
Hu, W.S., H.W. Chen, R.Y. Zhang, C.Y. Huang, and C.F. Shen. (2011). The expression levels of outer membrane proteins STM1530 and OmpD, which are influenced by the CpxAR and BaeSR two-component systems, play important roles in the ceftriaxone resistance of Salmonella enterica serovar Typhimurium. Antimicrob. Agents Chemother. 55: 3829-3837.
|
Jadhav, S.R., K.S. Rao, Y. Zheng, R.M. Garavito, and R.M. Worden. (2013). Voltage dependent closure of PorB class II porin from Neisseria meningitidis investigated using impedance spectroscopy in a tethered bilayer lipid membrane interface. J Colloid Interface Sci 390: 211-216.
|
Jasim, R., M.A. Baker, Y. Zhu, M. Han, E.K. Schneider-Futschik, M. Hussein, D. Hoyer, J. Li, and T. Velkov. (2018). A Comparative Study of Outer Membrane Proteome between Paired Colistin-Susceptible and Extremely Colistin-Resistant Klebsiella pneumoniae Strains. ACS Infect Dis. [Epub: Ahead of Print]
|
Jeanteur, D., J.H. Lakey, and F. Pattus. (1991). The bacterial porin superfamily: sequence alignment and structure prediction. Mol. Microbiol. 5: 2153-2164.
|
Jeanteur, D., J.H. Lakey, and F. Pattus. (1994). The porin superfamily: diversity and common features. In: Bacterial Cell Wall (J.M.Ghuysen and R. Hakenbeck, eds.). Elsevier, Amsterdam, pp. 363-380.
|
Jones, R.A., A.E. Jerse, and C.M. Tang. (2024). Gonococcal PorB: a multifaceted modulator of host immune responses. Trends Microbiol. 32: 355-364.
|
Kattner C., Zaucha J., Jaenecke F., Zachariae U. and Tanabe M. (2013). Identification of a cation transport pathway in Neisseria meningitidis PorB. Proteins. 81(5):830-40.
|
Kattner, C., S. Pfennig, P. Massari, and M. Tanabe. (2015). One-step purification and porin transport activity of the major outer membrane proteins P2 from Haemophilus influenzae, FomA from Fusobacterium nucleatum and PorB from Neisseria meningitidis. Appl Biochem Biotechnol 175: 2907-2915.
|
Kesireddy, A., K.R. Pothula, J. Lee, D.S. Patel, M. Pathania, B. van den Berg, W. Im, and U. Kleinekathöfer. (2019). Modeling of Specific Lipopolysaccharide Binding Sites on a Gram-Negative Porin. J Phys Chem B 123: 5700-5708.
|
Keyhani, N.O., X.B. Li, and S. Roseman. (2000). Chitin catabolism in the marine bacterium Vibrio furnissii. Identification and molecular cloning of a chitoporin. J. Biol. Chem. 275: 33068-33076.
|
Koebnik, R., K.P. Locher, and P. Van Gelder. (2000). Structure and function of bacterial outer membrane proteins: barrels in a nutshell. Mol. Microbiol. 37: 239-253.
|
Lee, S., N.G. Housden, S.A. Ionescu, M.H. Zimmer, R. Kaminska, C. Kleanthous, and H. Bayley. (2020). Transmembrane Epitope Delivery by Passive Protein Threading through the Pores of the OmpF Porin Trimer. J. Am. Chem. Soc. 142: 12157-12166.
|
Liko, I., M.T. Degiacomi, S. Lee, T.D. Newport, J. Gault, E. Reading, J.T.S. Hopper, N.G. Housden, P. White, M. Colledge, A. Sula, B.A. Wallace, C. Kleanthous, P.J. Stansfeld, H. Bayley, J.L.P. Benesch, T.M. Allison, and C.V. Robinson. (2018). Lipid binding attenuates channel closure of the outer membrane protein OmpF. Proc. Natl. Acad. Sci. USA 115: 6691-6696.
|
Lou, H., M. Chen, S.S. Black, S.R. Bushell, M. Ceccarelli, T. Mach, K. Beis, A.S. Low, V.A. Bamford, I.R. Booth, H. Bayley, and J.H. Naismith. (2011). Altered antibiotic transport in OmpC mutants isolated from a series of clinical strains of multi-drug resistant E. coli. PLoS One 6: e25825.
|
Lovelle, M., T. Mach, K.R. Mahendran, H. Weingart, M. Winterhalter, and P. Gameiro. (2011). Interaction of cephalosporins with outer membrane channels of Escherichia coli. Revealing binding by fluorescence quenching and ion conductance fluctuations. Phys Chem Chem Phys 13: 1521-1530.
|
Luo, X.-.W., P.-.L. Li, Y.-.J. Zhai, Y.-.S. Pan, G.-.Z. Hu, and D.-.D. He. (2024). Upregulation of outer membrane porin gene contributed to enhancement of azithromycin susceptibility in multidrug-resistant. Microbiol Spectr 12: e0391823.
|
Lv, T., F. Dai, Q. Zhuang, X. Zhao, Y. Shao, M. Guo, Z. Lv, C. Li, and W. Zhang. (2020). Outer membrane protein OmpU is related to iron balance in Vibrio alginolyticus. Microbiol Res 230: 126350.
|
Manoharan-Basil, S.S., Z. Gestels, S. Abdellati, E.A. Akomoneh, and C. Kenyon. (2023). Evidence of horizontal gene transfer within in 19 018 whole-genome spp. isolates: a global phylogenetic analysis. Microb Genom 9:.
|
Masi, M., M. Winterhalter, and J.M. Pagès. (2019). Outer Membrane Porins. Subcell Biochem 92: 79-123.
|
Massari, P., C.A. King, A.Y. Ho, and L.M. Wetzler. (2003). Neisserial PorB is translocated to the mitochondria of HeLa cells infected with Neisseria meningitidis and protects cells from apoptosis. Cell. Microbiol. 5: 99-109.
|
Mayse, L.A. and L. Movileanu. (2023). Gating of β-Barrel Protein Pores, Porins, and Channels: An Old Problem with New Facets. Int J Mol Sci 24:.
|
Milenkovic, S., J. Wang, S. Acosta-Gutierrez, M. Winterhalter, M. Ceccarelli, and I.V. Bodrenko. (2023). How the physical properties of bacterial porins match environmental conditions. Phys Chem Chem Phys 25: 12712-12722.
|
Nikaido, H. (1992). Porins and specific channels of bacterial outer membranes. Mol. Microbiol. 6: 435-442.
|
Novović, K., S. Mihajlović, M. Dinić, M. Malešević, M. Miljković, M. Kojić, and B. Jovčić. (2018). Acinetobacter spp. porin Omp33-36: Classification and transcriptional response to carbapenems and host cells. PLoS One 13: e0201608.
|
Pagel, M., V. Simonet, J. Li, M. Lallemand, B. Lauman, and A.H. Delcour. (2007). Phenotypic Characterization of Pore Mutants of the Vibrio cholerae Porin OmpU. J. Bacteriol. 189(23): 8593-8600.
|
Pal, A., L. Dhara, and A. Tripathi. (2019). Contribution of upregulation & OmpC/Ompk36 loss over the presence of towards carbapenem resistance development among pathogenic spp. Indian J Med Res 149: 528-538.
|
Park, H.J., S.W. Lee, and S.W. Han. (2014). Proteomic and functional analyses of a novel porin-like protein in Xanthomonas oryzae pv. oryzae. J Microbiol 52: 1030-1035.
|
Patel, D.S., S. Re, E.L. Wu, Y. Qi, P.E. Klebba, G. Widmalm, M.S. Yeom, Y. Sugita, and W. Im. (2016). Dynamics and Interactions of OmpF and LPS: Influence on Pore Accessibility and Ion Permeability. Biophys. J. 110: 930-938.
|
Pathania, M., S. Acosta-Gutierrez, S.P. Bhamidimarri, A. Baslé, M. Winterhalter, M. Ceccarelli, and B. van den Berg. (2018). Unusual Constriction Zones in the Major Porins OmpU and OmpT from Vibrio cholerae. Structure 26: 708-721.e4.
|
Pavez, M., C. Vieira, M.R. de Araujo, A. Cerda, L.M. de Almeida, N. Lincopan, and E.M. Mamizuka. (2016). Molecular mechanisms of membrane impermeability in clinical isolates of Enterobacteriaceae exposed to imipenem selective pressure. Int J Antimicrob Agents. [Epub: Ahead of Print]
|
Perini, D.A., A. Alcaraz, and M. Queralt-Martín. (2019). Lipid Headgroup Charge and Acyl Chain Composition Modulate Closure of Bacterial β-Barrel Channels. Int J Mol Sci 20:.
|
Peterson, J.H., L. Yang, J.C. Gumbart, and H.D. Bernstein. (2025). Conserved lipid-facing basic residues promote the insertion of the porin OmpC into the outer membrane. mBio 16: e0331924.
|
Prajapati, J.D., C.J.F. Solano, M. Winterhalter, and U. Kleinekathöfer. (2018). Enrofloxacin Permeation Pathways across the Porin OmpC. J Phys Chem B 122: 1417-1426.
|
Prilipov, A., P.S. Phale, R. Koebnik, C. Widmer, and J.P. Rosenbusch. (1998). Identification and characterization of two quiescent porin genes, nmpC and ompN, in Escherichia coli BE. J. Bacteriol. 180: 3388-3392.
|
Rassam, P., K.R. Long, R. Kaminska, D.J. Williams, G. Papadakos, C.G. Baumann, and C. Kleanthous. (2018). Intermembrane crosstalk drives inner-membrane protein organization in Escherichia coli. Nat Commun 9: 1082.
|
Reddy, B.L. and M.H. Saier, Jr. (2016). Properties and Phylogeny of 76 Families of Bacterial and Eukaryotic Organellar Outer Membrane Pore-Forming Proteins. PLoS One 11: e0152733.
|
Rodríguez-Morales, O., M. Fernández-Mora, I. Hernández-Lucas, A. Vázquez, J.L. Puente, and E. Calva. (2006). Salmonella enterica serovar Typhimurium ompS1 and ompS2 mutants are attenuated for virulence in mice. Infect. Immun. 74: 1398-1402.
|
Rumbo, C., M. Tomás, E. Fernández Moreira, N.C. Soares, M. Carvajal, E. Santillana, A. Beceiro, A. Romero, and G. Bou. (2014). The Acinetobacter baumannii Omp33-36 porin is a virulence factor that induces apoptosis and modulates autophagy in human cells. Infect. Immun. 82: 4666-4680.
|
Sajeev-Sheeja, A., E. Smorodina, and S. Zhang. (2023). Structural bioinformatics studies of bacterial outer membrane β-barrel transporters and their AlphaFold2 predicted water-soluble QTY variants. PLoS One 18: e0290360.
|
Santiviago, C.A., J.A. Fuentes, S.M. Bueno, A.N. Trombert, A.A. Hildago, L.T. Socias, P. Youderian, and G.C. Mora. (2002). The Salmonella enterica sv. Typhimurium smvA, yddG and ompD (porin) genes are required for the efficient efflux of methyl viologen. Mol. Microbiol. 46: 687-698.
|
Schulz, G.E. (1996). Porins: general to specific, native to engineered passive pores. Curr. Opin. Struc. Biol. 6: 485-490.
|
Simonet, V.C., A. Baslé, K.E. Klose, and A.H. Delcour. (2003). The Vibrio cholerae porins OmpU and OmpT have distinct channel properties. J. Biol. Chem. 278: 17539-17545.
|
Smani, Y., J. Dominguez-Herrera, and J. Pachón. (2013). Association of the outer membrane protein Omp33 with fitness and virulence of Acinetobacter baumannii. J Infect Dis 208: 1561-1570.
|
Solov''eva, T.F., N.M. Tischenko, V.A. Khomenko, O.Y. Portnyagina, N.Y. Kim, G.N. Likhatskaya, O.D. Novikova, and M.P. Isaeva. (2014). Study of effect of substitution of the penultimate amino acid residue on expression, structure, and functional properties of Yersinia pseudotuberculosis OmpY porin. Biochemistry (Mosc) 79: 694-705.
|
Solov'eva, T., G. Likhatskaya, V. Khomenko, K. Guzev, N. Kim, E. Bystritskaya, O. Novikova, A. Stenkova, A. Rakin, and M. Isaeva. (2017). The impact of length variations in the L2 loop on the structure and thermal stability of non-specific porins: The case of OmpCs from the Yersinia pseudotuberculosis complex. Biochim. Biophys. Acta. 1860: 515-525. [Epub: Ahead of Print]
|
Solov'eva, T.F., G.N. Likhatskaya, V.A. Khomenko, A.M. Stenkova, N.Y. Kim, O.Y. Portnyagina, O.D. Novikova, E.V. Trifonov, E.A. Nurminski, and M.P. Isaeva. (2011). A novel OmpY porin from Yersinia pseudotuberculosis: structure, channel-forming activity and trimer thermal stability. J Biomol Struct Dyn 28: 517-533.
|
Song, J., C.A. Minetti, M.S. Blake, and M. Colombini. (1998). Successful recovery of the normal electrophysiological properties of PorB (class 3) porin from Neisseria meningitidis after expression in Escherichia coli and renaturation. Biochim. Biophys. Acta. 1370: 289-298.
|
Song, W., H. Bajaj, C. Nasrallah, H. Jiang, M. Winterhalter, J.P. Colletier, and Y. Xu. (2015). Understanding Voltage Gating of Providencia stuartii Porins at Atomic Level. PLoS Comput Biol 11: e1004255.
|
Srinivasan, V.B., M. Venkataramaiah, A. Mondal, V. Vaidyanathan, T. Govil, and G. Rajamohan. (2012). Functional characterization of a novel outer membrane porin KpnO, regulated by PhoBR two-component system in Klebsiella pneumoniae NTUH-K2044. PLoS One 7: e41505.
|
Stein, C., O. Makarewicz, J.A. Bohnert, Y. Pfeifer, M. Kesselmeier, S. Hagel, and M.W. Pletz. (2015). Three Dimensional Checkerboard Synergy Analysis of Colistin, Meropenem, Tigecycline against Multidrug-Resistant Clinical Klebsiella pneumonia Isolates. PLoS One 10: e0126479.
|
Suginta, W., K.R. Mahendran, W. Chumjan, E. Hajjar, A. Schulte, M. Winterhalter, and H. Weingart. (2011). Molecular analysis of antimicrobial agent translocation through the membrane porin BpsOmp38 from an ultraresistant Burkholderia pseudomallei strain. Biochim. Biophys. Acta. 1808: 1552-1559.
|
Suginta, W., S. Sanram, A. Aunkham, M. Winterhalter, and A. Schulte. (2021). The C2 entity of chitosugars is crucial in molecular selectivity of the Vibrio campbellii chitoporin. J. Biol. Chem. 101350. [Epub: Ahead of Print]
|
Tanabe, M., C.M. Nimigean, and T.M. Iverson. (2010). Structural basis for solute transport, nucleotide regulation, and immunological recognition of Neisseria meningitidis PorB. Proc. Natl. Acad. Sci. USA 107: 6811-6816.
|
Tang, X., H. Wang, F. Liu, X. Sheng, J. Xing, and W. Zhan. (2019). Recombinant outer membrane protein T (OmpT) of Vibrio ichthyoenteri, a potential vaccine candidate for flounder (Paralichthys olivaceus). Microb. Pathog. 126: 185-192.
|
Tran, Q.T., K.R. Mahendran, E. Hajjar, M. Ceccarelli, A. Davin-Regli, M. Winterhalter, H. Weingart, and J.M. Pagès. (2010). Implication of porins in β-lactam resistance of Providencia stuartii. J. Biol. Chem. 285: 32273-32281.
|
Tsugawa, H., A. Ogawa, S. Takehara, M. Kimura, and Y. Okawa. (2008). Primary structure and function of a cytotoxic outer-membrane protein (ComP) of Plesiomonas shigelloides . FEMS Microbiol. Lett. 281: 10-16.
|
Utsunomia, C., C. Hori, K. Matsumoto, and S. Taguchi. (2017). Investigation of the Escherichia coli membrane transporters involved in the secretion of d-lactate-based oligomers by loss-of-function screening. J Biosci Bioeng 124: 635-640.
|
Vergalli, J., I.V. Bodrenko, M. Masi, L. Moynié, S. Acosta-Gutiérrez, J.H. Naismith, A. Davin-Regli, M. Ceccarelli, B. van den Berg, M. Winterhalter, and J.M. Pagès. (2019). Porins and small-molecule translocation across the outer membrane of Gram-negative bacteria. Nat. Rev. Microbiol. [Epub: Ahead of Print]
|
Vostrikova, O.P., M.P. Isaeva, G.N. Likhatskaya, O.D. Novikova, N.Y. Kim, V.A. Khomenko, and T.F. Solov'eva. (2013). OmpC-like porin from outer membrane of Yersinia enterocolitica: molecular structure and functional activity. Biochemistry (Mosc) 78: 496-504.
|
Wang, J., N. Fertig, and Y.L. Ying. (2019). Real-time monitoring β-lactam/β-lactamase inhibitor (BL/BLI) mixture towards the bacteria porin pathway at single molecule level. Anal Bioanal Chem 411: 4831-4837.
|
Wang, S.Y., J. Lauritz, J. Jass, and D.L. Milton. (2003). Role for the major outer-membrane protein from Vibrio anguillarum in bile resistance and biofilm formation. Microbiology 149: 1061-1071.
|
Ward, M.J., P.R. Lambden, and J.E. Heckels. (1992). Sequence analysis and relationships between meningococcal class 3 serotype proteins and porins from pathogenic and non-pathogenic Neisserial species. FEMS Microbiol. Lett. 73: 283-289.
|
Wassef M., Abdelhaleim M., AbdulRahman E. and Ghaith D. (2015). The Role of OmpK35, OmpK36 Porins, and Production of beta-Lactamases on Imipenem Susceptibility in Klebsiella pneumoniae Clinical Isolates, Cairo, Egypt. Microb Drug Resist. 21(6):577-80.
|
Whiting, G., C. Vipond, A. Facchetti, H. Chan, and J.X. Wheeler. (2019). Measurement of surface protein antigens, PorA and PorB, in Bexsero vaccine using quantitative mass spectrometry. Vaccine. [Epub: Ahead of Print]
|
Wise, M.G., E. Horvath, K. Young, D.F. Sahm, and K.M. Kazmierczak. (2018). Global survey of Klebsiella pneumoniae major porins from ertapenem non-susceptible isolates lacking carbapenemases. J. Med. Microbiol. 67: 289-295.
|
Yadav, S.K., J.K. Meena, M. Sharma, and A. Dixit. (2016). Recombinant outer membrane protein C of Aeromonas hydrophila elicits mixed immune response and generates agglutinating antibodies. Immunol Res 64: 1087-1099.
|
Yamamoto-Tamura, K., I. Kawagishi, N. Ogawa, and T. Fujii. (2015). A putative porin gene of Burkholderia sp. NK8 involved in chemotaxis toward β-ketoadipate. Biosci. Biotechnol. Biochem. 79: 926-936.
|
Ye, Y., L. Xu, Y. Han, Z. Chen, C. Liu, and L. Ming. (2018). Mechanism for carbapenem resistance of clinical Enterobacteriaceae isolates. Exp Ther Med 15: 1143-1149.
|
Zachariae, U., T. Klühspies, S. De, H. Engelhardt, and K. Zeth. (2006). High resolution crystal structures and molecular dynamics studies reveal substrate binding in the porin Omp32. J. Biol. Chem. 281: 7413-7420.
|
Zwama, M., A. Yamaguchi, and K. Nishino. (2019). Phylogenetic and functional characterisation of the multidrug efflux pump AcrB. Commun Biol 2: 340.
|
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.1.1 | OmpF general porin. OmpF can deliver peptides of >6 KDa (epitopes) including protamine, through the pore lumen from the periplasm to the outside (Housden et al., 2010; Ghale et al. 2014). For cephalosporin antibiotics, the interaction strength series is ceftriaxone > cefpirome > ceftazidime (Lovelle et al. 2011). An unfolded protein such as colicin E9 can thread through OmpF from the outside to reach the periplasm (Housden et al. 2013). Polynucleotides can pass through OmpF (Hadi-Alijanvand and Rouhani 2015). LPS influences the movement of bulk ions (K+ and
Cl-), but the ion selectivity of OmpF is mainly affected by bulk ion
concentrations (Patel et al. 2016). OMPs such as OmpF cluster into islands that restrict their lateral mobility, while IMPs generally diffuse throughout the cell. Rassam et al. 2018 demonstrated that when transient, energy-dependent transmembrane connections are formed, IMPs become subjugated by the inherent organisation of OMPs, and that such connections impact IMP function. They showed that while establishing a translocon for import, colicin ColE9 sequesters the IMPs of the proton motive force (PMF)-linked Tol-Pal complex into islands mirroring those of colicin-bound OMPs. Through this imposed organisation, the bacteriocin subverts the outer-membrane stabilizing role of Tol-Pal, blocking its recruitment to cell division sites and slowing membrane constriction. The ordering of IMPs by OMPs via an energised inter-membrane bridge represents an emerging functional paradigm in cell envelope biology (Rassam et al. 2018). Colicin E9 (ColE9) disordered regions exploit OmpF for direction-specific
binding, which ensures the constrained presentation of an activating
signal within the bacterial periplasm (Housden et al. 2018). Anionic lipid binding can prevent closure of OmpF channels, thereby increasing access of antibiotics that use porin-mediated pathways (Liko et al. 2018). OmpF may be the major route of D-lactate/D-3-hydroxybutyrate oligo-ester secretion (Utsunomia et al. 2017). Lipid Headgroup Charge and Acyl Chain Composition Modulate Closure of the channel (Perini et al. 2019). Piperacillin, tazobactam, ampicillin and sulbactam interact strongly with OmpF, and may be transported (Wang et al. 2019). Gating kinetics are governed by lipid characteristics so that each stage
of a sequential closure is different from the previous one, probably
because of intra- or intermonomeric rearrangements (Perini et al. 2019). OmpF transports fosfomycin (Golla et al. 2019) and bacteriocins into cells. Polypeptide transport/binding processes generate an essentially irreversible, hook-like assembly that constrains an import activating peptide epitope between two subunits of the OmpF trimer (Lee et al. 2020). Physical properties of bacterial porins (OmpF and OmpC) match environmental conditionsof induction (Milenkovic et al. 2023). Enrofloxacin caused blockage of ion current through OmpF, depending on the side of addition to the assymetic bilayer containing lipopolysaccharide and the transmembrane voltage applied (Donoghue et al. 2023). OmpF homologs are OmpK35 of Klebsiella pneumoniae and OmpE35 of Enterobacteria cloacae, and these porins transport ciprofloxacin (Acharya et al. 2024). OmpC and OmpF outer membrane porins of Escherichia coli and Salmonella enterica form bona fide amyloids (Belousov et al. 2023). Mechanisms of ciprofloxacin translocation through OmpF and other major diffusion channels of the ESKAPE pathogens Klebsiella pneumoniae and Enterobacter cloacae have been elucidated (Acharya et al. 2024). The presence of the enterobacterial common antigen (ECA) and capsular polysaccharides CPSs in the outer membrane (OM) enhances the rigidity and stability of the
OM while reducing the pore size of OmpF and increasing its cation
selectivity (Gao et al. 2025). | Proteobacteria | OmpF of E. coli (P02931) |
| |
| 1.B.1.1.10 | Putative porin | γ-Proteobacteria | Putative porin of Dickeya dadantii |
| |
| 1.B.1.1.11 | Porin OmpPst1. Transports carbapenem antibiotics imipenem will slow flux and meropenem with rapid flux in a reconsituted ion conductance system (Bajaj et al. 2012). | Proteobacteria | OmpPst1 of Providencia stuartii |
| |
| 1.B.1.1.12 | High conductance Omp35 (OmpK35; OmpF) porin. Expression levels are important for beta-lactam/cephalosporin resistance (Bornet et al. 2004). 95% identical to the Klebsiella pneumoniae orthologue, OmpK35 (Taherpour and Hashemi 2013). In K. pneumoniae, colistin-based combination therapy with a carbapenem and/or tigecycline
was associated with significantly decreased mortality rates due in part to synergistic induction of porins K35 and K36 (Stein et al. 2015). They influence imipenem susceptibility as well (Wassef et al. 2015). These two porins play roles in conferring carbapenem resistance in K. pneumoniae (Hamzaoui et al. 2018; Ye et al. 2018). Loss of a single porin (OmpK35 or OmpK36 (TC# 1.B.1.1.19) in Klebsiella pneumoniae is paired with reductions in capsule, increased LPS, and up-regulated transcription of compensatory porin genes, but loss of both porins resulted in an increase in capsule production. Loss of OmpK35 alone or dual porin loss was further associated with reduced oxidative burst by macrophages
and increased ability of the bacteria to survive phagocytic killing (Brunson et al. 2019). | Proteobacteria | Omp35 of Enterobacter (Aerobacter) aerogenes |
| |
| 1.B.1.1.13 | Omp36 (OmpC) porin of 375 aas (Dé et al. 2001; Bornet et al. 2004). Mutations affect beta-lactam and carbapenem (imipenem) sensitivity (Pavez et al. 2016). | Proteobacteria | Omp36 of Enterobacter (Aerobacter) aerogenes |
| |
| 1.B.1.1.14 | Major voltage-independent outer membrane porin, OmpH (Chevalier et al. 1993). A 3D model was obtained using in silico modeling. OmpH is probably a homotrimeric, 16 stranded, β-barrel porin involved in the non-specific transport of small, hydrophilic molecules, serving osmoregulatory functions (Ganguly et al. 2015). | Proteobacteria | OmpH of Pasteurella multocida |
| |
| 1.B.1.1.15 | OmpU porin (cation-selective; PK/PCl = 14; bile salt inducible) (low permeability to bile) (Simonet et al., 2003). OmpU influences sensitivities to β-lactam antibiotics and sodium deoxycholate induction of biofilm formation and growth on large sugars (Pagel et al., 2007). The effective pore radus is 0.55 nm which increases with acidic pH but decreases with increasing ionic strength (Duret and Delcour 2010). OmpU induces target animal cell death after it inserts into host mitochondrial membranes (Gupta et al. 2015). The high resolution structures of OmpT and OmpU, the two major porins in V. cholerae, have been determined, and both have unusual constrictions that create narrower barriers
for small-molecule permeation and change the internal electric fields of
the channels (Pathania et al. 2018). Vibrio cholerae OmpU activates dendritic cells via TLR2 and the NLRP3 inflammasome (Dhar et al. 2023). Four conserved domains in OmpU are linked with resistance to bile and
host-derived antimicrobial peptides. Mutant strains for these domains
exhibit differential susceptibility patterns to these and other
antimicrobials. A mutant strain in which the
four domains of the clinical allele were exchanged for those of a sensitive strain
exhibits a resistance profile closer to a porin deletion mutant.
Finally, using phenotypic microarrays, novel functions of
OmpU and their connection with allelic variability were uncovered (Grant et al. 2023). | Bacteria | OmpU of Vibrio cholerae |
| |
| 1.B.1.1.16 | OmpC of 367 aas (Vostrikova et al. 2013). | Proteobacteria | OmpC of Yersinia enterocolitica |
| |
| 1.B.1.1.17 | OmpF of 243 aas (Vostrikova et al. 2013). | Proteobacteria | OmpF of Yersinia enterocolitica |
| |
| 1.B.1.1.18 | Putative porin of 381 aas | Proteobacteria | PP of Klebsiella pneumoniae |
| |
| 1.B.1.1.19 | Outer membrane porin, KpnO, OmpCKP or OmpK36 of 367 aas. Loss causes increased drug (e.g., carbapenem and imipenem, but not colistin) resistance (Wassef et al. 2015, Jasim et al. 2018) decreased virulence and increased susceptibility to gastrointestinal stress (García-Sureda et al. 2011; Srinivasan et al. 2012). Expression is under PhoBR control. Porin deficiency is a widespread phenomenon, probably accounting for elevated ertapenem resistance (Wise et al. 2018). Loss of OmpK36 gives rise to carbapenem resistance (i.e., meropenem resistance) (Pal et al. 2019). | Proteobacteria | KpnO of Klebsiella pneumoniae |
| |
| 1.B.1.1.2 | PhoE phosphoporin. The 3-d structure is available (PDB#1PHO) | Bacteria | PhoE of E. coli |
| |
| 1.B.1.1.20 | Outer membrane porin 1, OmpPst1 or Omp-Pst1. Transports beta lactams with decreased efficiency in the order of ertapenem > cefepime > cefoxitin (Tran et al. 2010). 93% identical in sequence to 1.B.1.1.11. Voltage-gating of this porin and porin 2 (TC# 1.B.1.1.24) from the same organism have been analyzed (Song et al. 2015). Facing channels are open in any two adjacent porin structures, suggesting that dimers and trimers not only promote cell-to-cell contact but also contribute to intercellular communication (El-Khatib et al. 2018). | Proteobacteria | OmpPst1 of Providencia stuartii |
| |
| 1.B.1.1.21 | Outer membrane non-specific porin, OmpN or OmpS2, under the control of SoxS, and coregulated with the ydbK gene encoding pyruvate:flavodoxin oxidoreductase which plays a role in protection against oxidative stress (Prilipov et al. 1998; Fàbrega et al. 2012). MicC sRNA acts together with the σE envelope stress response pathway to control OmpN levels in response to β-lactam antibiotics (Dam et al. 2017). | Proteobacteria | OmpN of E. coli |
| |
| 1.B.1.1.22 | Outer membrane porin of 383 aas, OmpS2. Activated by OmpR and LeuO (Fernández-Mora et al. 2004). | Proteobacteria | OmpS2 of Salmonella typhi |
| |
| 1.B.1.1.23 | Outer membrane trimeric porin, OmpY (also called OmpN or OmpC2) of 360 aas. The effects of the length of loop L2 on function and stability have been studied (Solov'eva et al. 2017). | Proteobacteria | OmpY of Yersinia pseudotuberculosis |
| |
| 1.B.1.1.24 | Porin 2 (Omp-Pst2 or OmpPst2) of 365 aas. Voltage gating is observed for Omp-Pst2, where the binding of cations
in-between L3 and the barrel wall results in exposing a conserved
aromatic residue in the channel lumen, thereby halting ion permeation.
Comparison of Omp-Pst1 (TC# 1.B.1.1.20) with Omp-Pst2 suggested
that their differing sensitivities to voltage is encoded in the hydrogen-bonding
network anchoring L3 onto the barrel wall. The
strength of this network governs the probability of cations
binding behind L3. That Omp-Pst2 gating is observed only when ions flow
against the electrostatic potential gradient of the channel
suggests a possible role for this porin in the regulation of charge distribution across the outer membrane and bacterial homeostasis (Song et al. 2015). | Proteobacteria | OmpPst2 of Providenica stuartii |
| |
| 1.B.1.1.25 | Anion-selective, voltage-sensitive porin, VCA_1008, of 331 aas with a pore exclusion limit of 6.9 nm (Goulart et al. 2015). | | VCA_1008 of Vibrio cholerae |
| |
| 1.B.1.1.26 | The mature outer membrane protein, OmpC of 342 aas. Elicits an immune response (Yadav et al. 2016). | | OmpC of Aeromonas hydrophila |
| |
| 1.B.1.1.27 | Outer membrane porin, OmpS1 of 394 aas. mutants defective for OmpS1 are attenuated for virulence in mice (Rodríguez-Morales et al. 2006). | | OmpS1 in Salmonella enterica serovar Typhi |
| |
| 1.B.1.1.28 | OmpF of 364 aas and 1 N-terminal TMS. It has an abnormally high closing potential possibly due to charged residues and intramolecular bonds (Chistyulin et al. 2019). | | OmpF of Yersinia ruckeri |
| |
| 1.B.1.1.29 | Outer membrane porin, OmpE36, of 369 aas and 1 N-terminal TMS. It is 90% identical to Omp36 (TC# 1.B.1.1.13). Divalent cations, especially Ca2+, stabilize its binding to LPS molecules (Kesireddy et al. 2019). | | OmpE36 of Klebsiella aerogenes (Enterobacter aerogenes) |
| |
| 1.B.1.1.3 | OmpC general porin. Expression of OmpC and OmpF is reciprocally regulated by the EnvZ/OmpR sensor kinase/response regulator system (Egger et al. 1997). Mutants isolated from patients with MDR E. coli, resistant to several antibiotics, showed decreased permeability to these antibiotics (Lou et al. 2011). The diffusion route of the fluoroquinolone, enrofloxacin, through the OmpC porin has been reported (Prajapati et al. 2018). This system also transports azithromycin (Luo et al. 2024). Conserved lipid-facing basic residues promote the insertion of the porin OmpC into the E. coli outer membrane (Peterson et al. 2025). | Bacteria | OmpC of E. coli |
| |
| 1.B.1.1.30 | OmpU of 337 aas. OmpU is recognized by Toll-Like receptors in monocytes and macrophages for the induction of proinflammatory responses (Gulati et al. 2019). | | OmpU of Vibrio parahaemolyticus |
| |
| 1.B.1.1.31 | OmpC of 384 aas and 1 N-terminal TMS. | | OmpC of Enterobacter hormaechei |
| |
| 1.B.1.1.32 | Outer membrane porin, OmpN or OmpK37 of 374 aas and 1 N-terminal TMS. It was found to have higher mutability as a distinguishing feature which
makes it an important protein in monitoring the evolving resistances in
microorganisms | | OmpN of Klebsiella pneumoniae |
| |
| 1.B.1.1.33 | Putative porin of 467 aas | | Porin of Alteromonadales bacterium TW-7 |
| |
| 1.B.1.1.4 |
Weakly anion-selective NmpC (OmpD) porin (Prilipov et al. 1998). Transports methyl benzyl viologen, ceftriaxone and hydrogen peroxide in Salmonella species (Hu et al. 2011; Calderón et al. 2010). | Bacteria | NmpC of E. coli |
| |
| 1.B.1.1.5 | LC (lysogenic conversion) porin. Can replace OmpC and OmpF and is therefore probably non-selective (Fralick et al. 1990). Synthesis is subject to catabolite repression mediated by the cyclicAMP receptor protein, CRP (Blasband and Schnaitman 1987).
| Phage | LC porin of phage PA-2 |
| |
| 1.B.1.1.6 | Major outer membrane porin, OpnP. Probably orthologous to the E. coli OmpF. Expression of the opnP gene is activated by EnvZ and regulated by temperatur (Forst et al. 1995; Forst and Tabatabai 1997). | Bacteria | OpnP of Xenorhabdus nematophilus |
| |
| 1.B.1.1.7 | ComP porin. A virulence factor essential for cytotoxicity and apoptosis by this enteric pathogen (Tsugawa et al., 2008) | Bacteria | ComP of Plesiomonas shigelloides (A0JCJ5) |
| |
| 1.B.1.1.8 | Trimeric 16 TMS non-specific porin, Omp-EA (Elazer et al., 2007) | Bacteria | Omp-EA of Erwinia amylovora (A0RZH5) |
| |
| 1.B.1.1.9 | OmpU porin (weakly cation-selective; expression is induced by bile salts; OmpU mediates bile salt resistance) (Wang et al., 2003). This protein is 65% identical to OmpU of Vibrio alginolyticus which is involved in iron balance (Lv et al. 2020). | Bacteria | OmpU of Listonella (Vibrio) anguillarum (Q8GD13) |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.11.1 | Putative porin (based on homology) of 375 aas | Proteobacteria | Putative porin of Helicobacter hepaticus |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.2.1 | Omp25 of 255 aas. Associates with CarO (Siroy et al. 2005). | Proteobacteria | Omp25 of Acidetobacter baumannii |
| |
| 1.B.1.2.2 | Putative porin of 251 aas | Proteobacteria | PP of Shewanella piezotolerans |
| |
| 1.B.1.2.3 | Putative porin of 306 aas | Proteobacteria | PP of Colwellia psychrerythraea (Vibrio psychroerythus) |
| |
| 1.B.1.2.4 | Outer membrane porin, Omp33 or Omp33-36. This protein is a virulence factor and induces apoptosis in the host (Rumbo et al. 2014; Smani et al. 2013). It is involved in carbapenem resistance and is highly polymorphic (Novović et al. 2018). | Proteobacteria | Omp33-36 of Acinetobacter baumannii |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.3.1 | Omp2 porin. Several differing sequences for this protein can be found in GenBank. The one with acc# P46025 is 80% identical to the one listed here in TCDB. It may transport β-lactams and novobiocin (Zwama et al. 2019). | Bacteria | Omp2 of Haemophilus influenzae |
| |
| 1.B.1.3.2 | OmpP2 porin (transports NAD and NMN; transport Km=5 mM; may also serve as a general diffusion porin) (Andersen et al., 2003). Its solute transport activity with size exclusion limit has been described (Kattner et al. 2015). | Bacteria | OmpP2 of Haemophilus influenzae (Q48217) |
| |
| 1.B.1.3.3 | Putative porin | γ-Proteobacteria | Putative porin of Haemophilus parainfluenzae |
| |
| 1.B.1.3.4 | Putative porin | β-Proteobacteria | Putaive porin of Neisseria sp. |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.4.1 | Omp porin | Bacteria | Omp porin of Bordetella pertussis |
| |
| 1.B.1.4.2 | Phthalate porin, OphP (Chang et al. 2009).
| Bacteria | OphP of Burkholderia capacia (C0LZS0) |
| |
| 1.B.1.4.3 | Porin of 38 KDa,Omp38 in Burkholderia pseudomallei, the causative agent of
melioidosis, an infectious disease of animals and humans. MDR can be due to mutations in Omp38. Ion current
blockages of reconstituted Omp38 by seven antimicrobial agents occurred in a concentration-dependent
manner with the translocation on-rate following the order:
norfloxacin>ertapenem>ceftazidime>cefepime>imipenem>meropenem>penicillin
G (Suginta et al. 2011). Also allows transport of neutral sugars and numerous antimicrobial agents including cephalosporin and carbapenem (Aunkham et al. 2014). | Proteobacteria | Omp38 of Burkholderia pseudomallei |
| |
| 1.B.1.4.4 | Outer membrane porin of 353 aas (Brunen et al. 1991). | Proteobacteria | OMP of Acidovorax delafieldii |
| |
| 1.B.1.4.5 | Porin-like protein | Chlorobi | Porin of Chlorobium phaeobacteroides |
| |
| 1.B.1.4.6 | Putative porin of 248 aas | Proteobacteria | PP of Burkholderia cepacia (T0ET67) |
| |
| 1.B.1.4.7 | Outer membrane porin, OMPNK8 of 380 aas. Probably involved in transport of and chemotaxis toward β-ketoadipate; encoded by a gene (orf1) on a megaplasmid (pNK8) that carries the gene cluster (orf1-tfdT-CDEF), encoding chlorocatechol-degrading enzymes. orf1 is induced by the presence of 3-chlorobenzoate as is the tfd operon (Yamamoto-Tamura et al. 2015). | | OMPNK8 of Burkholderia sp. NK8 |
| |
| 1.B.1.4.8 | Outer membrane porin of 394 aas and 16 predicted beta strands, isolated from an endosymbiont of a trypanosomatid protozoan (Andrade et al. 2011). | | Porin of an endosymbiont of Crithidia deanei |
| |
| 1.B.1.4.9 | OmpQ porin of 364 aas | | OmpQ of Bordetella parapertussis |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.5.1 | Oma1 porin (Class 1) (Tanabe et al., 2010) | Bacteria | Oma1 of Neisseria gonorrhoeae |
| |
| 1.B.1.5.2 | PorA porin, cation selective at pH > 6; anion selective at pH < 4 (a continuum electrodiffusion model accounts for the results) (Cervera et al., 2008). Both PorA and PorB have been used for vaccine development (Whiting et al. 2019). | Bacteria | PorA of Neisseria meningitidis |
| |
| 1.B.1.5.3 | Major outer membrane protein IB (OMB) (slightly cation-selective porin) | Bacteria | OMB of Neisseria sicca |
| |
| 1.B.1.5.4 |
PorB porin (Song et al. 1998; Tanabe et al., 2010). The 2.3 Å structure has been determined by x-ray
crystallography. There are three putative solute translocation pathways
through the channel pore: One pathway transports anions nonselectively,
one tranports cations nonselectively, and one facilitates the specific
uptake of sugars (Kattner et al. 2012). Regulated by ATP binding (Tanabe et al., 2010). Exhibits voltage-dependent closure (Jadhav et al. 2013). Its unique solute transport activity with size exclusion limit has been described (Kattner et al. 2015). The β-lactam antibiotic ampicillin binds to PorB (Bartsch et al. 2019). Recombination in loop regions between pathogenic and non-pathogenic Neisseria spp. has been observed, suggested a mechanism for developing variation in drug resistance (Manoharan-Basil et al. 2023). Gonococcal PorB is a multifaceted modulator of host immune responses (Jones et al. 2024). | Bacteria | PorB porin of Neisseria meningitidis |
| |
| 1.B.1.5.6 | Porin of 434 aas amd 1 N-terminal TMS. | Acidobacteria | Porin of Holophaga foetida |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.6.1 | Anion-selective porin protein 32, Omp32. The structure is known to 1.5 Å resolution (Zachariae et al. 2006). | Bacteria | Porin protein 32 of Comamonas (Delftia) acidovorans |
| |
| 1.B.1.6.2 | Outer membrane porin of 304 aas (Brunen et al. 1991). | Proteobacteria | OMP of Acidovorax delafieldii |
| |
| 1.B.1.6.3 | Outer membrane porin of 319 aas (Brunen et al. 1991). | Proteobacteria | OMP of Acidovorax delafieldii |
| |
| 1.B.1.6.4 | Outer membrane porin of 313 aas | Chrysiogenetes | Porin of Desulfurispirillum indicum |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.7.1 | Chitoporin, ChiP of 366 aas and 1 N-terminal TMS. Its synthesis is induced by (GlcNAcn, n = 2-6, but not by GlcNAc or other sugars. A nulll mutant did not grow on GlcNAc3 and transported a nonmetabolizable analogue of GlcNAc2 at a reduced rate. (Keyhani et al., 2000). | Bacteria | ChiP of Vibrio furnissii |
| |
| 1.B.1.7.2 | Sugar-specific chitoporin of 375 aas, ChiP. The best substrate is chitohexose, but ChiP transports a variety of chitooligosaccharides. Trp136 is important for the
binding affinity for
chitohexaose (Chumjan et al. 2015). X-ray crystal structures of ChiP from V. harveyi in the presence and
absence of chito-oligosaccharides have been solved (Aunkham et al. 2018). Structures without bound sugar reveal
a trimeric assembly with an unprecedented closing of the transport pore
by the N-terminus of a neighboring subunit. Substrate binding ejects
the pore plug to open the transport channel.The structures explain the exceptional affinity of ChiP for
chito-oligosaccharides and point to an important role of the N-terminal
gate in substrate transport (Aunkham et al. 2018). Hydrogen-bonds contribute to sugar permeation (Chumjan et al. 2019). This protein is 90% identical to the chitoporin of the Vibrio campbellii chitoporin (Aunkham et al. 2020). The C2 entity of chitosugars is crucial for the molecular selectivity of the Vibrio campbellii chitoporin (Suginta et al. 2021). | Proteobacteria | ChiP of Vibrio harveyi |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.8.1 | Low ion selective porin (PK/PCl = 4), OmpT (high permeability to bile) (Simonet et al., 2003). OmpT has an effective radius of 0.43nm, and acidic pH, high ionic strength, or exposure to polyethyleneglycol stabilizes a less conductive state (Duret & Delcour, 2010). It binds the biofilm matrix protein, Bap1, which influences antimicrobial peptide (polymyxin B and LL-37) resistance (Duperthuy et al. 2013). The high resolution structures of OmpT and OmpU, the two major porins in V. cholerae, have been determined, and both have unusual constrictions that create narrower barriers
for small-molecule permeation and change the internal electric fields of
the channels (Pathania et al. 2018). | Bacteria | OmpT of Vibrio cholerae (AAC28105) |
| |
| 1.B.1.8.2 | Putative uncharacterized protein | None | Tresu_2327 of Treponema succinifaciens |
| |
| 1.B.1.8.3 | Porin-like protein H (37 kDa outer membrane protein) | None | ompH of Photobacterium profundum ) |
| |
| 1.B.1.8.4 | OmpT of 322 aas and 1 N-terminal TMS. OmpT is a promising vaccine candidate against V. ichthyoenteri infections in fish (Tang et al. 2019). | | OmpT of Vibrio ichthyoenteri |
| |
| 1.B.1.8.5 | OmpH, porin-like protein H precursor of 298 aas and 1 N-terminal TMS | | OmpH of Reinekea blandensis |
| |
| 1.B.1.8.6 | Porin of 352 aas | | Porin of Vibrio mediterranei |
| |
| Examples: |
| TC# | Name | Organismal Type | Example |
| 1.B.1.9.1 | The outer membrane porin, M35 (Easton et al., 2005) | Bacteria | M35 of Moraxella catarrhalis (AAX99225) |
| |
| 1.B.1.9.2 | Major porin of 369 aas, involved in anaerobic respiration, positively regulated by both CRP and FNR, OmpS38 or Omp35 (Gao et al. 2015). | | OmpS38 of Shewanella oneidensis |
| |
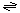 Solute (in)
Solute (in)