1.C.10 The Pore-forming Haemolysin E (HlyE) Family
The HlyE family consists of a single protein, HlyE, and its close orthologues from species of Escherichia, Shigella and Salmonella. The E. coli protein is a functionally well characterized, pore-forming chromosomally-encoded haemolysin also called ClyA (cytolysin A), Hpr, and silent haemolysin, SheA. It consists of 303 aas (34 kDa). Its transcription is positively controlled by SlyA, a regulator found in several enteric bacteria. HlyE forms stable, moderately cation-selective transmembrane pores with a diameter of 2.5-3.0 nm in lipid bilayers. The protein binds cholesterol, and pore formation in a membrane is stimulated if the membrane contains cholesterol. It forms oligomeric assemblies in the membrane (Wai et al., 2003). Common and divergent structural and functional traits that distinguish the various ClyA family PFTs have been reviewed (Bräuning and Groll 2018). ClyA, an alpha toxin with its inserted wedge shaped bundle of inserted alpha helices, induces asymmetry across the membrane leaflets in comparison with alpha hemolysin (αHL), a beta toxin (Varadarajan et al. 2020).
The crystal structure of soluble E. coli HlyE has been solved to 2.0 Å resolution, and visualization of the lipid-associated form of the toxin at low resolution has been achieved by electron microscopy. The structure is different from other toxins, exhibiting an elaborate helical bundle some 100 Å long. It oligomerizes in the presence of lipid transmembrane to form predominantly octameric or dodecameric pores (Tzokov et al., 2006; Ludwig et al., 2010). The complexes are conformationally variable. The pores are longer than expected from the dimensions of the soluble protein suggesting that conformational changes occur on pore formation (Tzokov et al., 2006). HlyE protomers retain an α-helical structure when oligomerized to form a pore consisting of parallel HlyE protomers (Hunt et al., 2008).
The HlyE cytotoxin has recently been shown to be exported from the bacterium and assembled in vesicles derived from the outer membrane of E. coli (Wai et al., 2003). It forms oligomeric pore assemblies in the outer membrane, and the toxic activity of this outer membrane assembled toxin towards animal cells proved to be higher than that of the purified protein. Thus, outer bacterial membrane vesicles contribute to the activation and delivery of toxins (Wai et al., 2003).
The soluble monomer of ClyA must undergo large conformational changes to form the transmembrane pore. Mueller et al., 2009 reported the 3.3 A crystal structure of the 400 kDa dodecameric transmembrane pore formed by ClyA (HlyE). The tertiary structure of ClyA protomers in the pore is substantially different from that in the soluble monomer. The conversion involves more than half of all residues. It results in large rearrangements, up to 140 A, of parts of the monomer, reorganization of the hydrophobic core, and transitions of beta-sheets and loop regions to alpha-helices. The large extent of interdependent conformational changes indicates a sequential mechanism for membrane insertion and pore formation.
ClyA, an α-pore forming toxin from pathogenic Escherichia coli (E. coli) and Salmonella enterica, assembles into an oligomeric structure in solution in the absence of either bilayer membranes or detergents at physiological temperature (Fahie et al. 2013). These oligomers can rearrange to create transmembrane pores when in contact with detergents or biological membranes. Intrinsic fluorescence measurements revealed that oligomers adopted an intermediate state found during the transition between monomer and transmembrane pore, suggesting that the water-soluble oligomer represents a prepore intermediate state. Moreover, ClyA does not form transmembrane pores on E. coli lipid membranes. Because ClyA is delivered to the target host cell in an oligomeric conformation within outer membrane vesicles (OMVs), ClyA apparently forms a prepore oligomeric structure independently of the lipid membrane within the OMV, a non-classical pathway to attack eukaryotic host cells (Fahie et al. 2013).
The ClyA monomer possesses an α-helical bundle with a β-sheet subdomain (the beta-tongue) previously believed to be critical for pore assembly and/or insertion. Oligomerization of ClyA pores transforms the beta-tongue into a helix-turn-helix that embeds into the lipid bilayer. Fahie et al. 2018 showed that mutations of the β-tongue did not prevent oligomerization or transmembrane insertion, but substitution mutants yielded pores with decreased conductance while a deletion mutation resulted in pores that rapidly closed following membrane association. Our results suggest that the beta-tongue plays a structural role in stabilizing the open conformation of the transmembrane domain.
The generalized transport reaction catalyzed by HlyE is:
Small molecules (in) 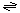 small molecules (out)
small molecules (out)