1.A.65 The Coronavirus Viroporin E Protein (Viroporin E) Family
Viroporins are a growing family of viral proteins able to enhance membrane permeability, promoting virus budding. The viroporin activity of the E protein from murine hepatitis virus (MHV), a member of the coronaviruses, resulted in exit of labeled nucleotides from E. coli cells to the cytoplasm upon expression of MHV E. In addition, enhanced entry of the antibiotic hygromycin B occurred at levels comparable to those observed with the viroporin 6K from Sindbis virus. Mammalian cells are also readily permeabilized by the expression of the MHV E protein. Finally, brefeldin A powerfully blocks the viroporin activity of the E protein in BHK cells, suggesting that an intact vesicular system is necessary for this coronavirus to permeabilize mammalian cells (Madan et al., 2005). The E protein has been described as a cation-selective Ca2+ channel (Harrison et al. 2022). The E channel maintains its multi-ionic non-specific neutral character even in concentrated solutions of Ca2+. In contrast to previous studies, Surya et al. 2023 found no evidence that SARS-CoV-2 E channel activation requires a particular voltage, high calcium concentrations or low pH, in agreement with available data from SARS-CoV-1 E. In addition, sedimentation velocity experiments suggest that the E channel population is mostly pentameric, but very dynamic and probably heterogeneous, consistent with the broad distribution of conductance values typically found in electrophysiological experiments. The latter has been explained by the presence of proteolipidic channel structures (Surya et al. 2023).
More recently, coronavirus (CoV) envelope (E) protein ion channel activity was determined in channels formed in planar lipid bilayers by peptides representing either the transmembrane domain of severe acute respiratory syndrome CoV (SARS-CoV) E protein, or the full-length E protein. Both of them formed voltage-independent ion-conductive pores with symmetric ion transport properties (Verdiá-Báguena et al., 2012). Mutations N15A and V25F located in the transmembrane domain prevented ion conductivity. E protein derived channels showed no cation preference in non-charged lipid membranes, whereas they behaved as pores with mild cation selectivity in negatively-charged lipid membranes. Thus, the ion conductance was controlled by the lipid composition of the membrane. Lipid charge also regulated the selectivity of a HCoV-229E E protein derived peptide. These results suggested that the lipids are functionally involved in E protein ion channel activity, forming a protein-lipid pore (Verdiá-Báguena et al. 2013). Refinement of the e-protein structure in a native-like environment by molecular dynamics simulations has been achieved, and it shows that it induced local membrane curvature while decreasing local lipid order (Yang et al. 2022). The SARS-CoV-2 envelope protein forms clustered pentamers in lipid bilayers (Somberg et al. 2022).
HydroDock can build hydrated drug-target complexes from scratch. The program requires only the dry target and drug structures and produces their complexes with appropriately positioned water molecules. As a test application of the protocol, Zsidó et al. 2021 built the structures of amantadine derivatives in complex with the influenza M2 transmembrane ion channel. The repositioning of amantadine derivatives from this influenza target to the SARS-CoV-2 envelope protein was also investigated. Excellent agreement was observed between experiments and the structures determined by HydroDock. The atomic resolution complex structures showed that water plays a similar role in the binding of amphipathic amantadine derivatives to transmembrane ion channels of both influenza A and SARS-CoV-2. While the hydrophobic regions of the channels capture the bulky hydrocarbon group of the ligand, the surrounding waters direct its orientation parallel with the axes of the channels via bridging interactions with the ionic ligand head (Zsidó et al. 2021). Mechanisms of host-membrane permeabilization by viroporins has been reviewed (Alcaraz and Nieva 2025).
The generalized reaction catalyzed by the MHV E protein is:
small molecules (out) 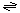 small molecules (in)
small molecules (in)