1.B.25 The Outer Membrane Porin (Opr) Family
The Opr family includes a number of porins in Pseudomonas species and other Gram-negative bacteria that appear to exhibit a variety of substrate selectivities. Thus, OprD2 of P. aeruginosa is specific for cationic amino acids, peptides and antibiotics, while PhaK is specific for phenolic compounds. GusC of E. coli is a 'membrane accessory protein' that facilitates uptake of glucuronides via the GusB permease. Many other specificities have been assigned (Tamber et al., 2006).
Tamber et al. (2006) have characterized several of the 19 paralogues in Pseudomonas aeruginosa. They fall into two fairly closely related phylogenetic clusters. Members of one, including OprD, exhibit specificities for amino acids and their derivatives while the other, including PhaK, are specific for organic acids. Another group of functionally characterized porins, determined by Tamber et al. (2006), are described in the table of the Opr family homologues by these authors.
In P. aeruginosa, the majority of small molecules are taken up by members of the OprD outer membrane protein family. Eren et al. (2012) showed that OprD channels require a carboxyl group in the substrate for efficient transport. They have renamed the family Occ, for outer membrane carboxylate channels. Occ channels can be divided into two subfamilies, based on their very different substrate specificities.
Liu et al. (2012) reported that the OccK proteins exhibit fairly distinct unitary conductance values including low (~40-100 pS) and medium (~100-380 pS) conductances. These proteins showed diverse single-channel dynamics of current gating transitions, revealing one (OccK3), two (OccK4, OccK5 and OccK6) and three (OccK1, OccK2 and OccK7) open sub-state kinetics with functionally distinct conformations. Anion selectivity is a conserved trait among the members of the OccK subfamily, confirming the presence of a net pool of positively charged residues within their central constriction.
The generalized transport reaction catalyzed by members of the Opr family is:
Substrate (out) 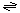 substrate (periplasm)
substrate (periplasm)